Each of these tools has pitfalls that can lead to false positives or false negatives and some tools are limited by minimum sequence length or are only geared to detect a limited range of virus families.īeyond discovery and detection, de novo annotation of contigs representing viruses presents a number of challenges. Strategies include detection of hallmark genes conserved within known virus families (but absent in cellular genomes) 4, 5, detection of short nucleotide sequences believed to be enriched in viruses 6 (or other machine learning approaches 7, 8), or the ratio of genes common to virus genomes versus genes common to non-viral sequences 9. Several tools have been developed to detect virus sequences in complex datasets. 
Sequence space thus covers at, at best, 0.0001% of the virosphere.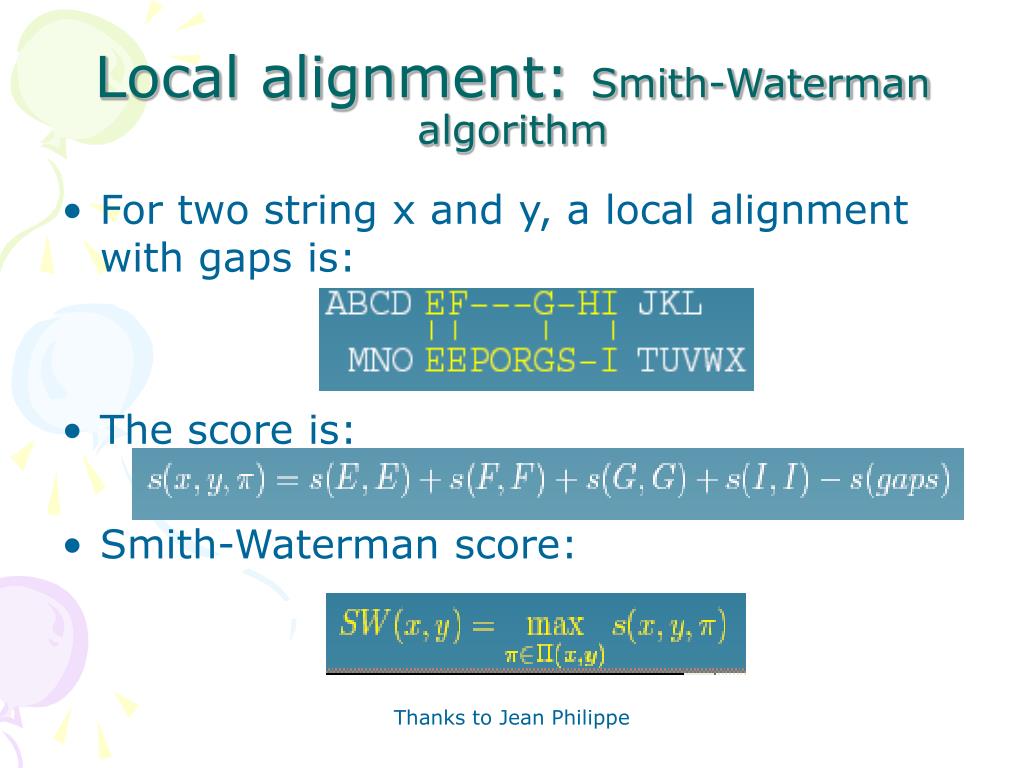 
Finally, at least hundreds of millions of virus species are likely to exist on Earth 2, but sequences for only tens of thousands of virus species are deposited in the central GenBank virus database and fewer than 10,000 virus species exist in the authoritative RefSeq database 3. Further, there are no universal genes found in all viral genomes that could be used to probe complex datasets for viruses, whereas cellular genomes can be detected through PCR targeting ribosomal genes and alignment of sequences to other single-copy marker genes 1. For example, animals and bacteria share homologous genes with more amino acid identity than even the most-conserved genes in some virus families (for example, GenBank sequences: polyomavirus Large T antigen and 60S ribosomal protein L23 ). Virus hunters have a challenging signal-to-noise problem to consider.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
